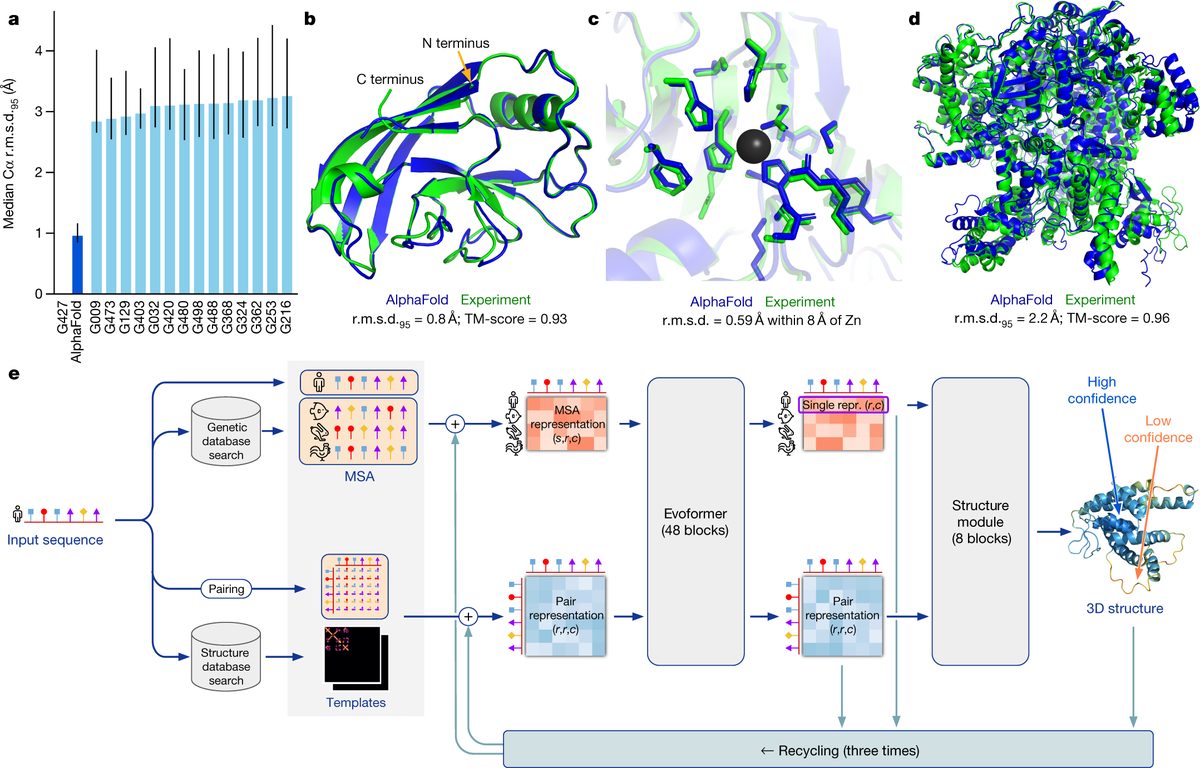
Determining the three-dimensional shape of a single protein used to take a graduate student an entire career. In March 2026, Google DeepMind released AlphaFold 4, a system that predicts the structure of multi-protein complexes in minutes rather than hours or days, extending a streak of capability gains that began when the original AlphaFold won a protein-prediction competition in 2018. Weeks earlier, DeepMind's drug-discovery spinoff Isomorphic Labs disclosed a proprietary model called IsoDDE that more than doubles AlphaFold 3's accuracy on key drug-design benchmarks, while a team at the National University of Singapore published D-I-TASSER, a tool that outperforms both AlphaFold 2 and AlphaFold 3 on difficult multi-domain proteins.
The practical stakes are enormous. Drug development currently averages more than a decade and roughly two billion dollars per approved medicine, with structure determination representing a critical bottleneck in identifying how candidate molecules bind to disease targets. AI systems that predict these interactions in seconds rather than months could compress early-stage discovery timelines dramatically. But the field is splitting: AlphaFold versions have historically been shared openly, while Isomorphic Labs' most powerful model is proprietary, sparking debate over whether the tools that reshape medicine should remain accessible to all researchers or become competitive advantages for well-funded companies.